2026.5·Bring your data, bring your team
All releases · May 4, 2026 · Jan Hellemans
This is the release where Clarida starts to feel like home for users coming from qbase+. Bring your existing experiments across without losing data. Tag your samples the way you have always wanted to. And, for the first time, invite your team into shared workspaces and organizations so the work no longer lives on a single person's account.
The qPCR field has been waiting a long time for a successor to qbase+. With 2026.5 we are closing the gap on the things people relied on every day, while building the collaborative foundation that desktop software could never offer.
Welcome aboard.
Jan
New features
Bring your qbase+ experiments to Clarida
Migrating from one analysis tool to another usually means leaving years of work behind. With 2026.5 you can import your qbase+ experiments directly into Clarida and pick up where you left off. Sample annotations, assay definitions, plate layouts, run results, exclusion decisions, and analysis settings all come across.
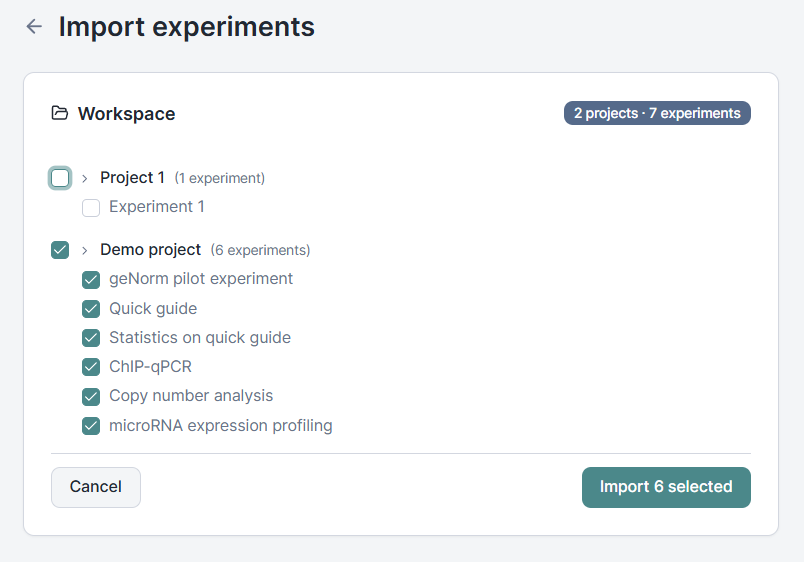
Imported experiments open in a read-only mode that protects the integrity of your historical data while keeping everything you need usable: review the analysis, annotate samples, regenerate reports, and export results. The structural pieces are locked, but the science is fully available. When you want to build new work on top, duplicate the experiment and make it your own.
Custom sample properties, all the way through to your charts
Tag your samples with the metadata that actually matters: tissue type, treatment, timepoint, donor, dilution, anything you need. Properties are typed, so a numeric concentration sorts the way you would expect and a categorical group filters cleanly.
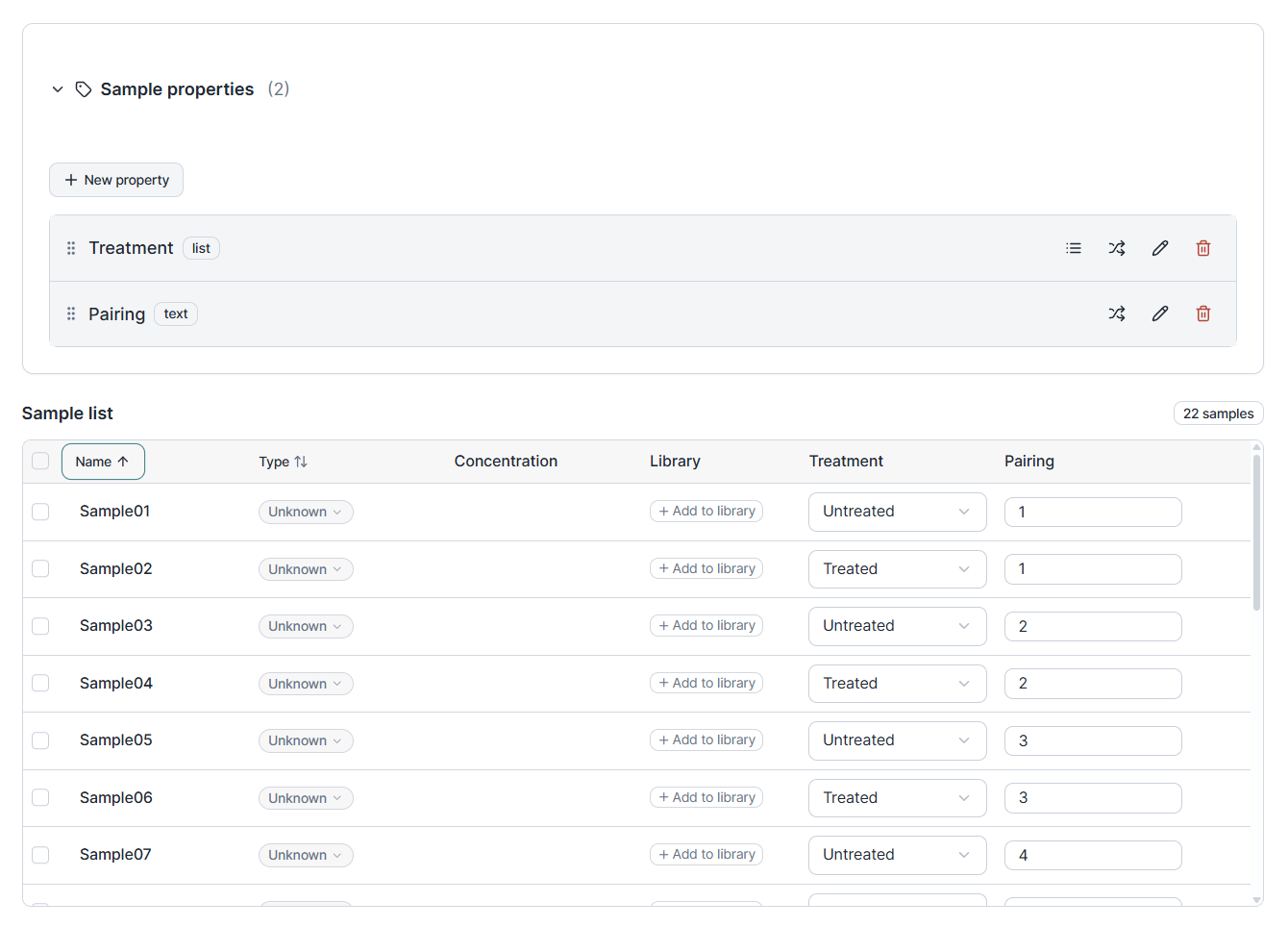
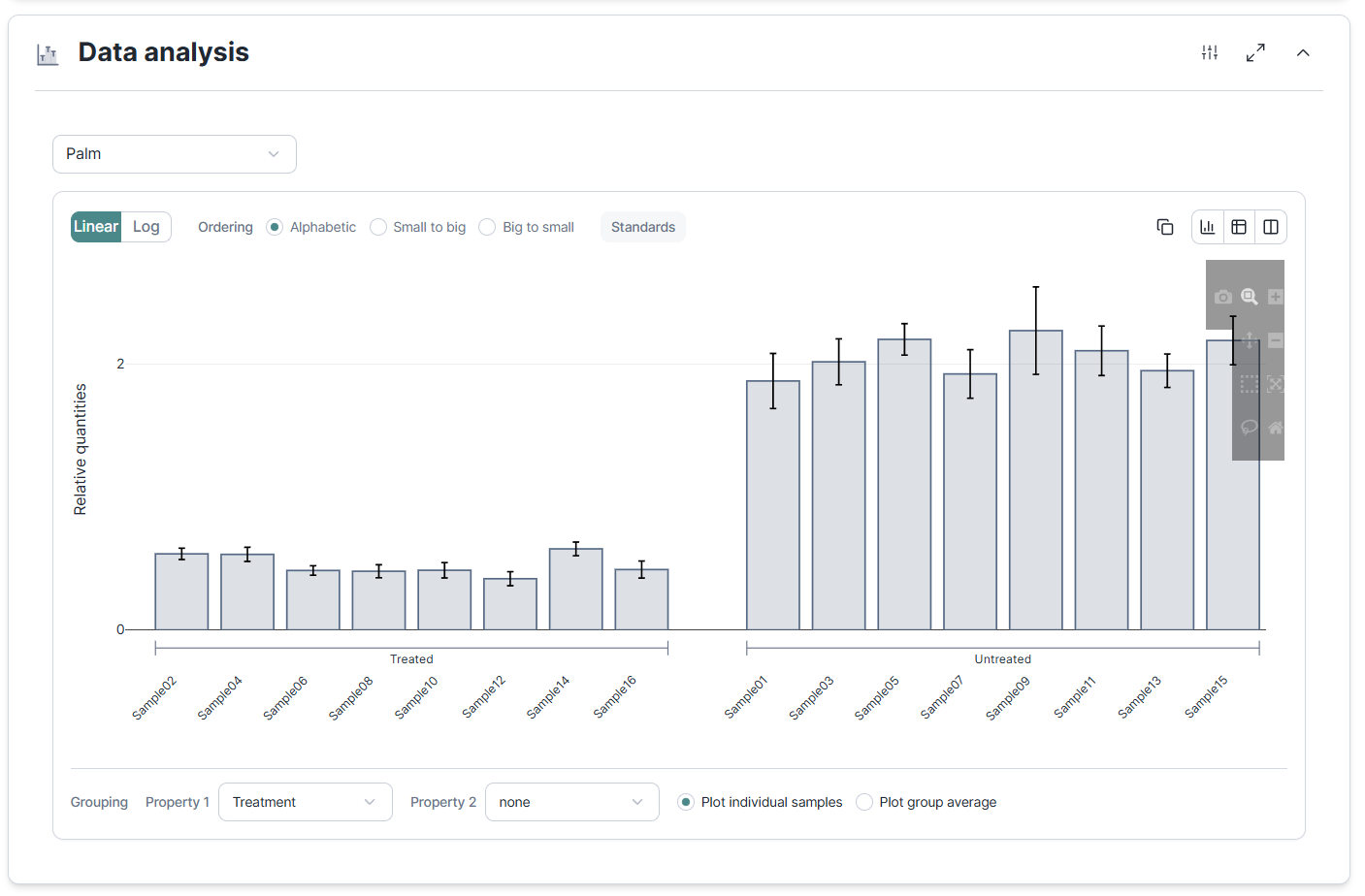
The properties do not stop at the table. In Data Analysis you can now sort and group your relative quantification results by any property you defined. Rather than reading a long list of sample names, see your treatment groups side by side, your timepoints in order, or your replicates collapsed by donor. The visualization adapts to how your experiment is actually organized.
Per-run cycling protocols and instrument selection
Cycling protocols and instrument choice now belong to the run, not the experiment. That matters when you run the same assay on two different instruments, or when one experiment spans a handful of runs with slightly different conditions. Each run carries its own protocol and instrument, and the MIQE traceability follows.
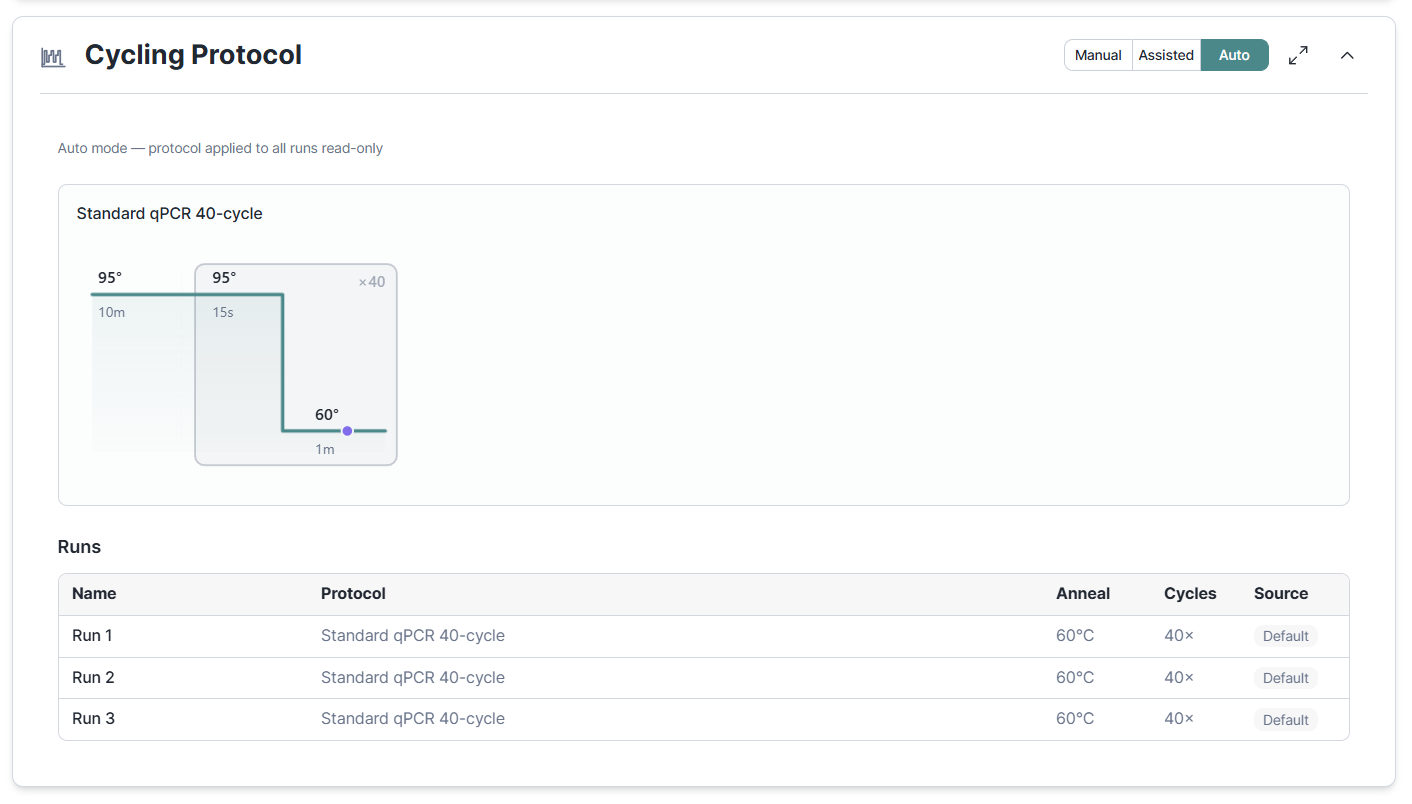
Previously, the design workflow derived the plate format for generating a run layout from the instrument set in the Cycling Protocol section. This logic has now been optimized with plate format being a setting of the Run Layout section and the Cycling Protocol section moved to the end so that run-specif protocols can be configured in response to the layout generated in the preivous step.
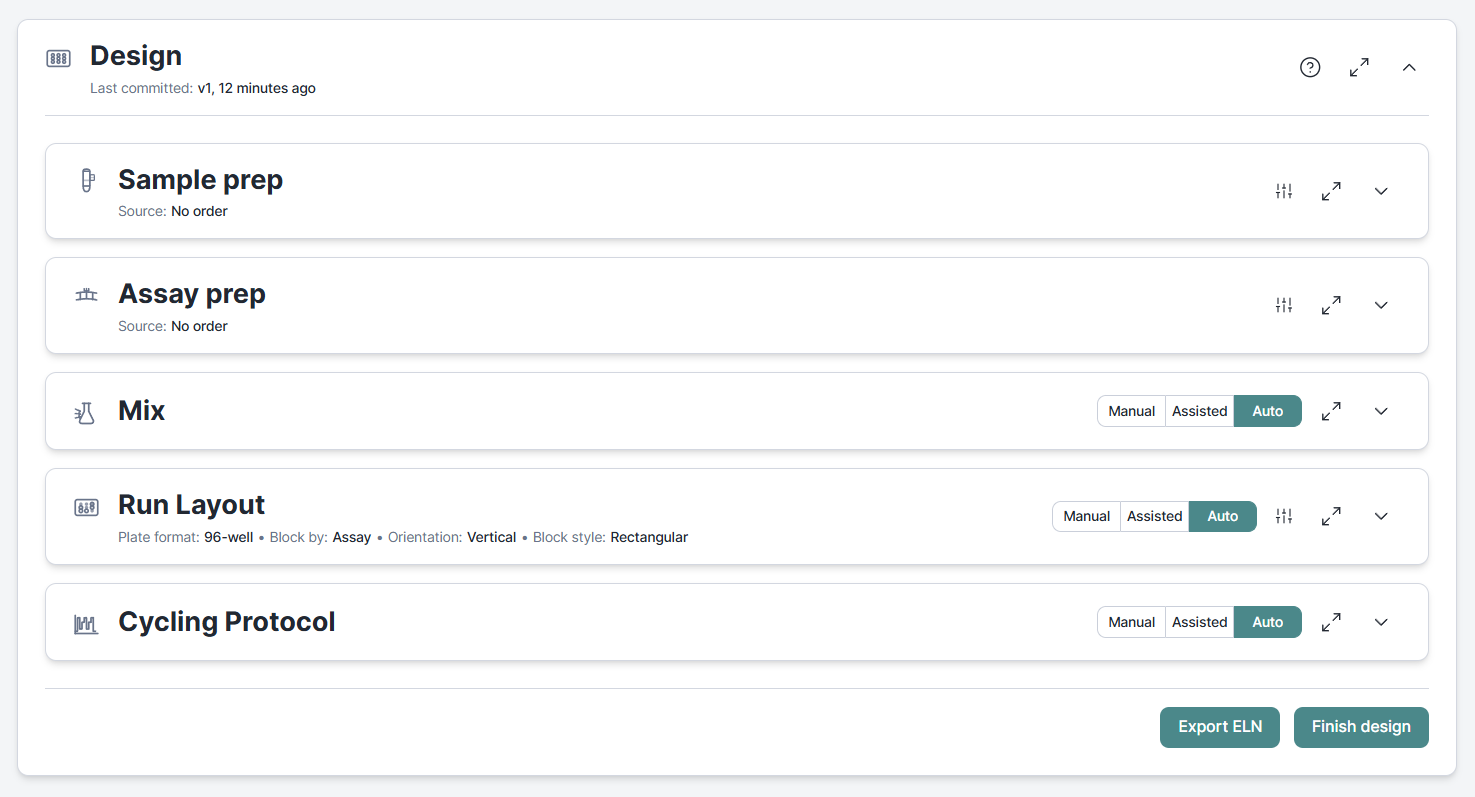
Run layouts based on source positions
In March we introduced auto-generated run-layouts with user control on the structure. This works fine if your assays and samples are stored in individual tubes and their relative positions do not matter. If your samples are stored in strips or a plate, or if you want to control the sample order, your only option was to take full control by manual run annotation. We now introduce Source positions that give you control on relative positions while also offering all the convenience of auto-generated layouts.
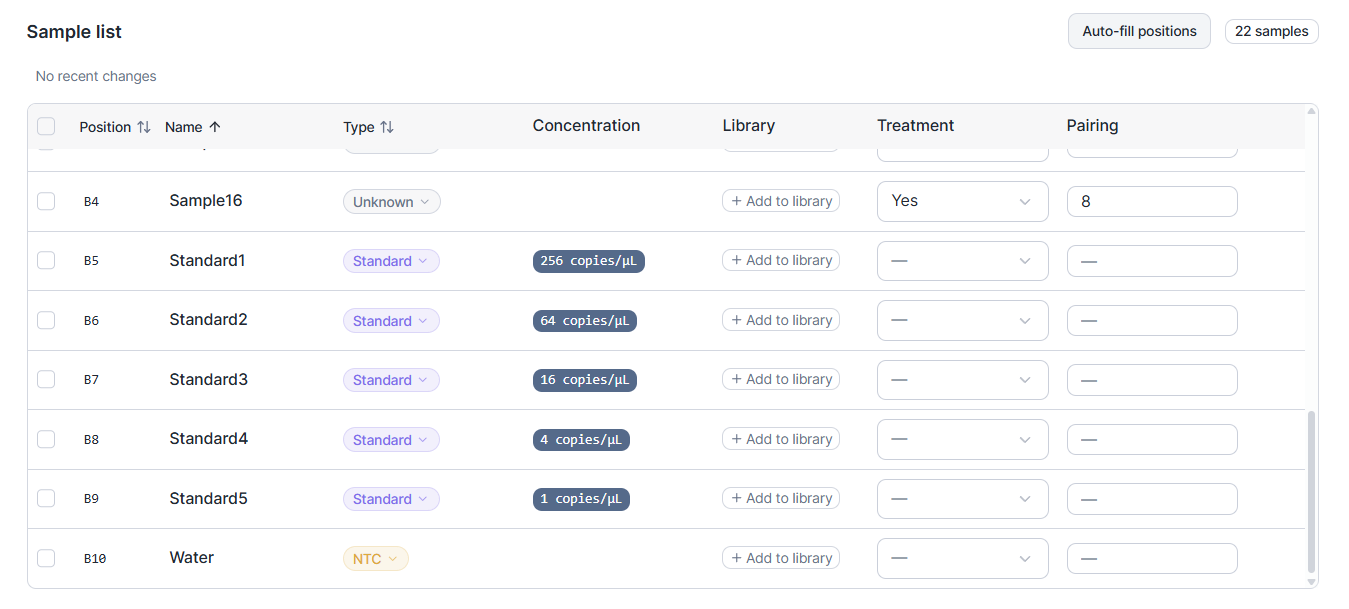
The result is a pipetting plan that matches what you can actually do at the bench, not a layout that looks tidy on screen but precludes time savings through the use of multi-channel pipets.
Storage Manager: one place for everything
Experiments, library samples, library assays, and tool outputs now live under a single Storage hub. Browse what you have, search across types, and open any item without losing your place in the rest of the app.
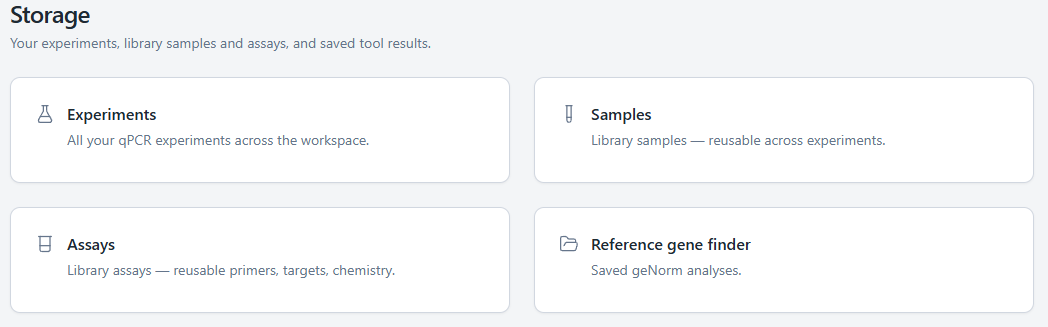
This sets the stage for the cross-experiment features we are building next.
Workspaces and organizations: bring your team in
Until now, a Clarida account was a one-person thing. With 2026.5 you can create a workspace, invite your colleagues, and share the work. Approve people who want to join with their institutional email, manage their roles, and remove access when someone leaves the lab. New users get a one-click sign-in link in their invite email, no passwords to set up before they can start.
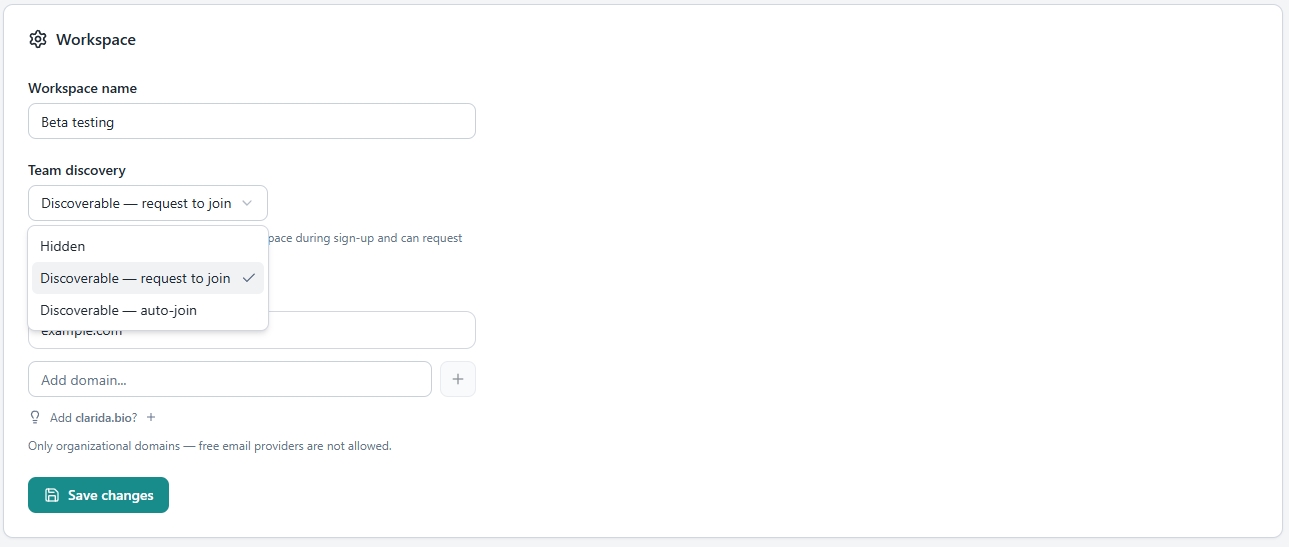
For larger setups — teaching labs, core facilities, partnerships — there is a layer above: organizations. An organization owns a shared usage pool that several workspaces can draw from, so a course coordinator can provision twenty student workspaces from one account, and a PI can give their lab access without per-seat fragmentation. Experiment duplication makes onboarding new team members straightforward: hand them a fully populated example and let them riff.
Freemium accounts stay single-workspace, paid plans unlock collaboration. The free tier still gets everything an individual researcher needs.
Quality of life
- Run Layout improvements · 384-well quadrant filling matches what robots and multi-channel pipets actually do, multi-plate spillover handles experiments larger than a single plate, and the plate visualization is improved to always reflect the current configuration.
- Reference Gene Finder PDF by email · Run an analysis on the public Reference Gene Finder and get a polished PDF emailed to you. No account needed for the report itself; an optional Clarida signup keeps your work for later.
- NRQ ordering and grouping · Grouping in Data Analysis can now respect your custom sample properties so you can present results the way your experiment is structured. Ordering of data by NRQ allows you to quickly sort samples by their expression level.
- Sample and assay volumes shown in Collect Material · See exactly how much of each you need before you start pipetting.
- Aside settings on every section · Every workflow section with settings now has a settings button in the header to quickly jump to those settings in the aside panel. When jumping between sections, clicking anywhere in a sections now activates it so the corresponding settings are properly synced.
- Concentration popover stays open · The unit dropdown in the concentration popover no longer dismisses the popover when opened.
- Standard sample updates retrigger downstream calculation · Change a standard sample and efficiency, normalization, and quantification all recalculate automatically.
- Feedback attachments · Attach more file types to feedback reports, and your contact information now auto-populates.
- Terms of service and privacy links on login and emails · Login page and invite/approval emails now link to the terms of service and privacy policy.
