Clarida early access is live
All posts · · Ward Hellemans
A while ago, we wrote about the PCR workflow problem - how the science has matured but the tools around it have not kept pace.
Today, we are inviting you in.
What we are building
Clarida is the operating system for quantitative biology. This first release starts with qPCR, where we have deep expertise and where the workflow pain is clear.
LIMS fights you. Excel forgets. Clarida connects.
If you have worked with LIMS or rigid lab systems, you know the tradeoff: powerful for compliance, but they fight you the moment your experiment does not fit their template. They assume they know your workflow better than you do.
If you have built your own Excel system (and let us be honest, most of us have), you know that tradeoff too: total flexibility, but no traceability. Good luck explaining what you did six months later.
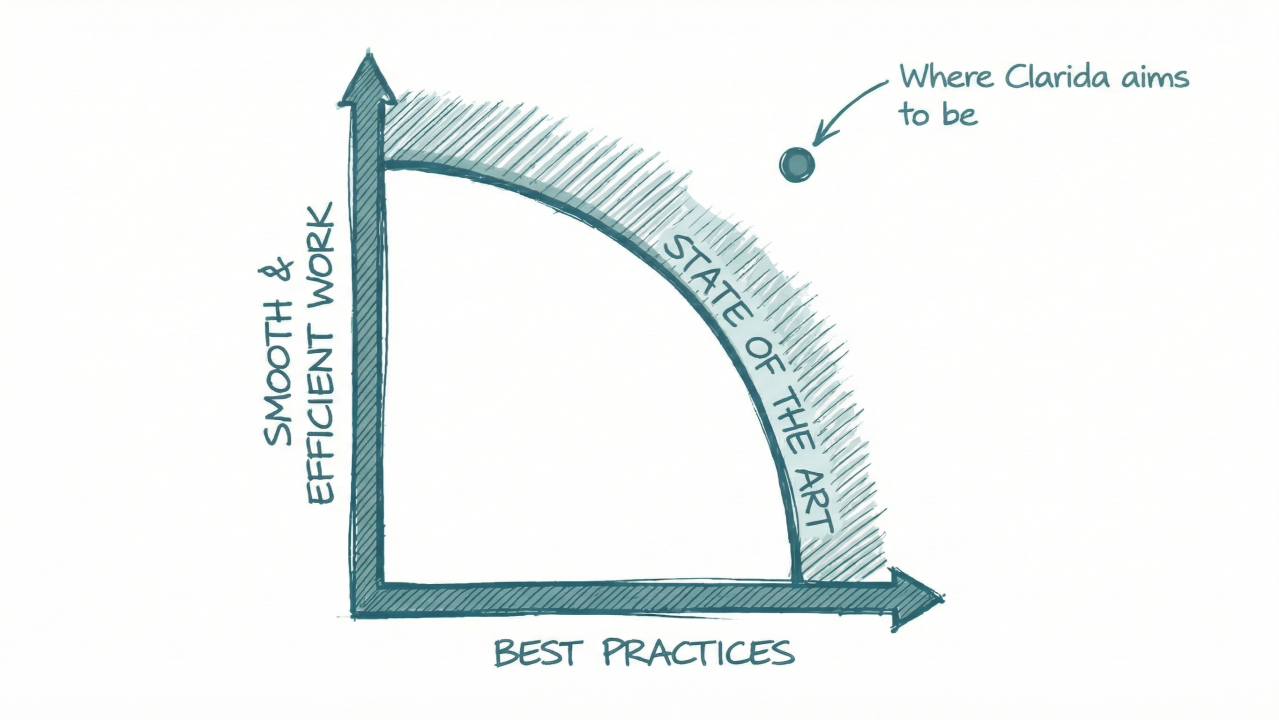
Most tools force a trade-off: efficiency or best practices. You pick a position on that line.
We built Clarida to move the line. Your data stays open and accessible, in formats you control, but the connections, the context, the audit trail are built in.
Open by design
Most scientific software locks you in. Proprietary formats, closed ecosystems, data you cannot easily export.
Clarida aims to become the home for your entire workflow, replacing the scattered Excel files, the disconnected tools, the fragmented process. But you stay because it works, not because you are trapped.
Your data stays open, exportable, in formats you control. Always.
Why we built this
Scientists are not failing at reproducibility because they do not care. They care deeply.
They are failing because they are forced to chase it across Excel sheets, vendor software, shared drives, and memory. Bits and pieces. Scattered tools. No connective tissue.
Clarida is where it finally comes together.
The D-DEAR framework
We have organized Clarida around the full arc of an experiment, from research question to actionable report. We call this framework D-DEAR.
Define - what are you trying to learn and why. Design - how you will structure the experiment. Execute - step-by-step wet-lab protocols with pre-calculated volumes and contextual checks. Analyze - coming soon. Report - coming soon.
If you know MIQE, think of D-DEAR as the workflow layer that makes MIQE compliance natural. MIQE tells you what to report. D-DEAR is about everything that happens before you report, structured so the reporting writes itself.
What is available now
This early access release covers the first three phases: Define, Design, and Execute.
We started here deliberately. This is where workflow chaos begins, and where errors cascade through everything downstream.
These phases are foundational in this release. Far from being fully fleshed out in width or depth, but functional. Enough to start working with, enough to show where we are headed.
The qPCR experiment workflow walks you through: define your project with research goals, background context, and reference links. Design experiments by configuring samples, assays, and mixing protocols. Execute with step-by-step protocols and Excel file export. Export to structured Excel workbooks, ready for your ELN.
Standalone tools for everyday lab work: Concentration Converter for switching between mass, molar, and copy number with built-in templates for organisms and dPCR platforms. Plate Homogeneity for detecting edge effects, spatial bias, CV and uniformity metrics. Lab Workbooks for downloading ready-made Excel templates or adding structure to your existing files.
Everything exports to Excel. No proprietary formats. No lock-in.
Coming next: Analyze and Report phases, dPCR experiment workflow, Reference gene finder, plus more width and depth to Define, Design, and Execute.
Try it
This is early access. Real, functional, but not finished. It will evolve based on how you actually use it.
That is the point.
We are looking for scientists willing to test, break things, and tell us what is missing. There is a feedback button built into the app, so you can flag issues or share ideas without leaving your workflow. Every piece of feedback helps us iterate faster.
Early access is free. Try it, break it, tell us.
Key takeaways
- Clarida early access is live at app.clarida.bio - free to use.
- Covers Define, Design, and Execute phases of the qPCR workflow.
- Includes standalone tools: Concentration Converter, Plate Homogeneity, and Lab Workbooks.
- Open by design: everything exports to Excel, no proprietary formats, no lock-in.
- Analyze and Report phases, dPCR workflow, and Reference gene finder coming next.
Ward Hellemans
Co-founder, Clarida
Former product manager for qbase+, where he led development of the commercial qPCR analysis platform. Now building Clarida to solve the full qPCR workflow problem — from experiment design to publication.
